
Surprisingly, neither STRIDE nor DSSP could identify a few π-helices that we had previously observed from graphical analysis of three-dimensional structures ( Al-Karadaghi et al., 1994, 1997).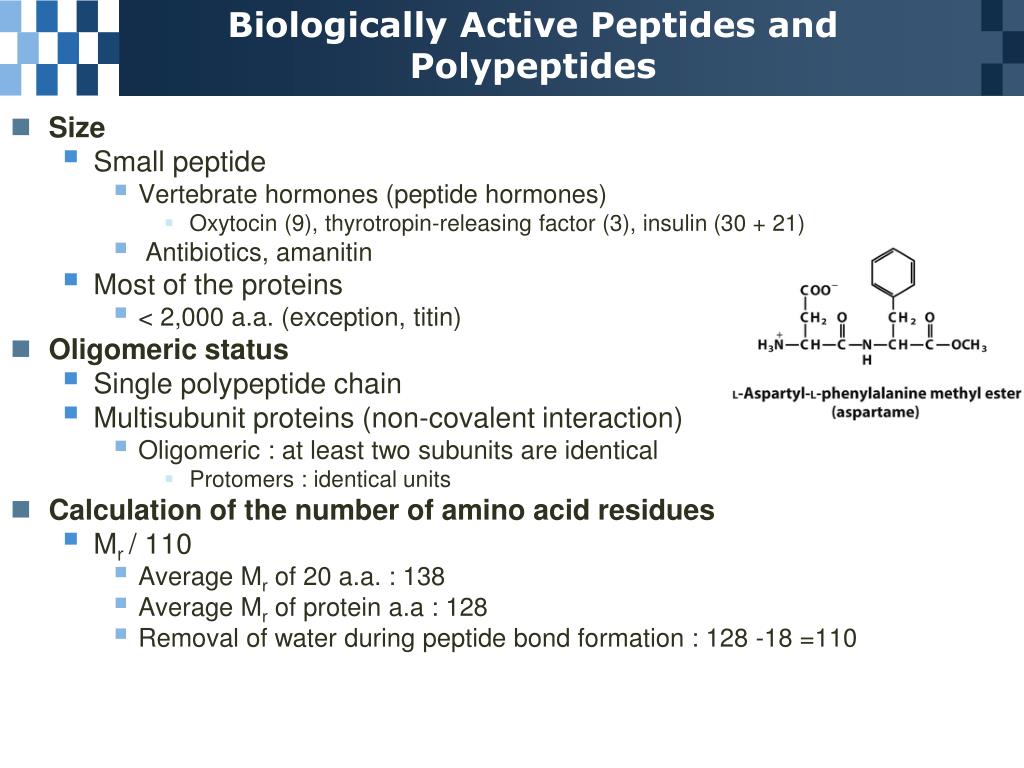 
In a similar search using STRIDE, we could not identify any additional hits (unpublished results). In a recent study, by using DSSP, only nine unique π-helices could be identified in a database of more than 6000 proteins ( Weaver, 2000). While DSSP identifies helices based on the presence of repeating ( i← i + n) hydrogen bonds with n corresponding to 3, 4 and 5 for 3 10-, α- and π-helices, respectively, STRIDE makes use of both hydrogen bonds and main chain dihedral angles to define secondary structures. Among these, DSSP ( Kabsch and Sander, 1983) and STRIDE ( Frishman and Argos, 1995) are the most widely used. Several algorithms have been designed to assign secondary structure automatically based on three-dimensional coordinates ( Kabsch and Sander, 1983 Richards and Kundrot, 1988 Frishman and Argos, 1995 Labesse et al., 1997). This suggests that the π-helix is not as unstable as previously believed. A few researchers have, however, found π-helices to be formed during molecular dynamics simulations of peptides ( Kovacs et al., 1995 Gibbs et al., 1997 Shirley and Brooks, 1997) with some reports of a transition from α-helix to π-helix structure ( Duneau et al., 1996 Lee et al., 2000). The rarity of the π-helix has been attributed to its instability due to the following properties: (1) the dihedral angles ϕ and Ψ are unfavorable, lying at the very edge of an allowed minimum energy region of the Ramachandran plot ( Ramachandran and Sasisekharan, 1968) (2) the larger radius of the π-helix means that main chain atoms are no longer in van der Waals contact along the helix axis, resulting in a hole too small for a water molecule to fill ( Low and Baybutt, 1952) (3) a large entropic cost is required to form a helix in which five residues need to be aligned to permit the ( i← i + 5) hydrogen bond ( Rohl and Doig, 1996). The π-helix, however, appears to be extremely rare. The α-helix is considered to be the most abundant form of secondary structure, accounting for about 31% of amino acid secondary structure states, while the 3 10-helix accounts for about 4% ( Baker and Hubbard, 1984 Barlow and Thornton, 1988). 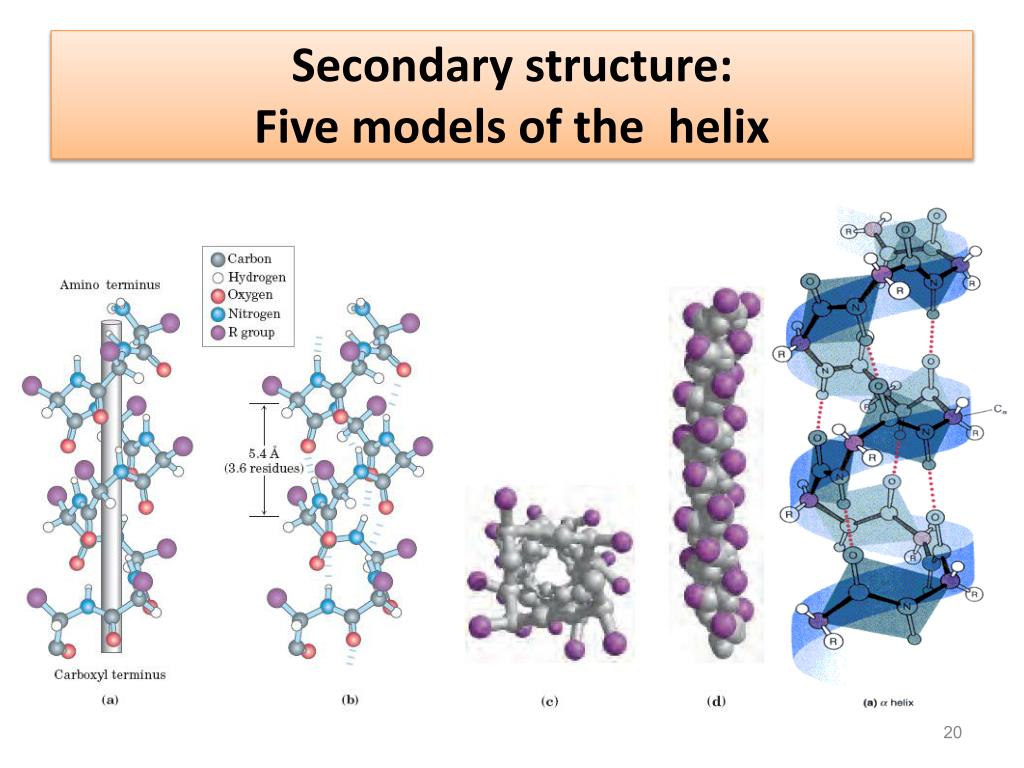
Of all the hypothesized helix types, only the α-helix, 3 10-helix and the π-helix have been observed in protein structures. The main stabilizing factor for helical structures in polypeptides is repeating hydrogen bonds between main chain carbonyl oxygen (C=O) and amide hydrogen (NH) groups with the α-helix characterized by an ( i← i + 4) pattern, the 3 10- and the π-helix by repeating ( i← i + 3) and ( i← i + 5) hydrogen bonds, respectively. Low and Baybutt also suggested the possibility of the 4.4 16-helix or π-helix ( Low and Baybutt, 1952). Donohue later considered the possibility of other types of helices (2.2 7, 3 10, 4.3 14 and 4.4 16) ( Donohue, 1953). Pauling and Corey first hypothesized the α-helix (3.6 13) and the γ-helix (5.1 17) structures ( Pauling and Corey, 1951). Helix types are usually designated as x y based on the number of residues per turn ( x) and the number of atoms in the ring closed by a hydrogen bond ( y) ( Donohue, 1953). Helices are a major type of secondary structure element found in proteins. Thus, we describe cases where the side chains of functionally important residues at every fourth position within a π-helix could be aligned and brought close together in a way that would not be allowed by any other helix type. The π-helix may have some functional advantages over other helical structures. In addition to hydrogen bonds, several other factors contribute to the stability of π-helices. Features that may contribute to the stability of the π-helical structure have also been identified. 
It also has distinctive amino acid preferences and it is conserved within functionally related proteins. This enabled us to show for the first time that the π-helix has structural parameters that are different from the hypothesized model values. Using an improved π-helix definition algorithm to search a non-redundant subset of high-resolution and well-refined protein structures, we found that almost every tenth protein contained a π-helix. Existing secondary structure definition methods find very few within the Protein Data Bank. The π-helix appears to be extremely rare and is considered to be unstable. The most abundant helix type in proteins is the α-helix, accounting for about 31% of amino acid secondary structure states, while the 3 10-helix accounts for about 4%.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
